9.A.15 The Autophagy-related Phagophore-formation Transporter (APT) Family
Autophagy is the degradation of a cell's own components within lysosomes (or the analogous yeast vacuole), and its malfunction contributes to a variety of human diseases. Atg9 is the sole integral membrane protein required in formation of the initial sequestering compartment, the phagophore, and it plays a key role in membrane transport as a phospholipid scramblase (Nakao and Nakano 2022); the phagophore presumably expands by vesicular addition to form a complete autophagosome. Atg9 functions at the phagophore assembly site (PAS) (Munakata and Klionsky, 2010). He et al. (2008) reported that Atg9 molecules self-associate independently of other known autophagy proteins in both nutrient-rich and starvation conditions. Mutational analyses revealed that self-interaction is critical for anterograde transport of Atg9 to the PAS. The ability of Atg9 to self-interact is required for both selective and nonselective autophagy at the step of phagophore expansion at the PAS. Atg9 multimerization facilitates membrane flow to the PAS for phagophore formation. It includes 6 putative TMSs.
Eukaryotic cells employ autophagy to degrade damaged or obsolete organelles and proteins (Umemiya et al., 2007). Central to this process is the formation of autophagosomes, double-membrane vesicles responsible for delivering cytoplasmic material to lysosomes. In the past decade many autophagy-related genes, Atg, have been identified that are required for autophagic functions. In all types of autophagy, a core molecular machinery has a critical role in forming sequestering vesicles, the autophagosome, which is the hallmark morphological feature of this dynamic process. Additional components allow autophagy to adapt to the changing needs of the cell (Xie and Klionsky, 2007). The phagophore-lysosome/vacuole fusion strategy can rescue defective autophagy due to failed phagophore closure in both yeast and Homo (Liang 2023). V-9302 in combination with trametinib inhibits the trametinib-induced autophagy, thereby enhancing pyroptosis in cancer cells (Liu et al. 2025).
In yeast, approximately 31 autophagy-related (Atg) proteins have been identified. Most of them reside at the phagophore assembly site (PAS). Geng et al. (2008) reported the application of fluorescence microscopy to study the amount of Atg proteins at the PAS. They found that an increase in the amount of Atg11 at the PAS enhanced the recruitment of Atg8 and Atg9 to this site and facilitated the formation of more cytoplasm-to-vacuole targeting vesicles. In response to autophagy induction, the amount of most Atg proteins remained unchanged at the PAS, whereas an enhanced recruitment of Atg8 and 9 at this site was observed. During autophagy, the amount of Atg8 at the PAS showed a periodic change, indicating the formation of autophagosomes, and both Atg8 and Atg12 were ubiquitinylated (Geng and Klionsky, 2008). Novel Atg proteins and the stages of their action have been identified (Backues et al. 2015).
Atg9 is the only characterized transmembrane protein that is absolutely required for Cvt vesicle formation, and it is proposed to carry membrane from peripheral donor sites to the phagophore assembly site where the vesicle forms. It is also a lipid scramblase that mediates autophagosomal membrane expansion (Matoba et al. 2020). Additional proteins, including Atg11, Atg23, and Atg27, are involved in the anterograde movement, whereas Atg1-Atg13 and Atg2-Atg18 are required for the retrograde return to the peripheral sites (Munakata and Klionsky, 2010). Atg11 and Atg23 show low sequence similarity to MLP1 and MLP2 of the Nuclear Pore Complex (NPC; 1.A.75.1.1). These proteins include repeat sequences. Atg1 is a serine, thereonine protein kinase; also called autophagy protein 3 or cytoplasm to vacuole targeting protein 10.
Intestinal Paneth cells limit bacterial invasion by secreting antimicrobial proteins, including lysozyme. However, invasive pathogens can disrupt the Golgi apparatus, interfering with secretion and compromising intestinal antimicrobial defense. Bel et al. 2017 showed that during bacterial infection, lysozyme is rerouted via secretory autophagy, an autophagy-based alternative secretion pathway. Secretory autophagy was triggered in Paneth cells by bacteria-induced endoplasmic reticulum (ER) stress, required extrinsic signals from innate lymphoid cells, and limited bacterial dissemination. Secretory autophagy was disrupted in Paneth cells of mice harboring a mutation in autophagy gene Atg16L1 that confers increased risk for Crohn's disease in humans. These findings identify a role for secretory autophagy in intestinal defense and suggest why Crohn's disease is associated with genetic mutations that affect both the ER stress response and autophagy (Bel et al. 2017).
The lysosome (or vacuole in yeast) is the central organelle responsible for cellular degradation and nutrient storage. Lysosomes receive cargo from the secretory, endocytic, and autophagy pathways. Many of these proteins and lipids are delivered to the lysosome membrane, and some are degraded in the lysosome lumen, whereas others appear to be recycled. Suzuki and Emr 2018 identified the transmembrane autophagy protein Atg27 as a physiological cargo recycled from the vacuole. Atg27 is delivered to the vacuole membrane and then recycled using a two-step recycling process. First, Atg27 is recycled from the vacuole to the endosome via the Snx4 complex and then from the endosome to the Golgi via the retromer complex TC# 9.A.3). During the process of vacuole-to-endosome retrograde trafficking, Snx4 complexes assemble on the vacuolar surface and recognize specific residues in the cytoplasmic tail of Atg27. This pathway maintains the normal composition and function of the vacuole membrane (Suzuki and Emr 2018).
Autophagy is a highly conserved recycling process of eukaryotic cells that degrades protein aggregates and damaged organelles with the participation of autophagy-related proteins. Membrane bending is a key step in autophagosome membrane formation and nucleation. A variety of autophagy- related proteins (ATGs) are needed to sense and generate membrane curvature, which then complete the membrane remodeling process. The Atg1 complex, Atg2-Atg18 complex, Vps34 complex, Atg12-Atg5 conjugation system, Atg8-phosphatidylethanolamine conjugation system, and transmembrane protein Atg9 promote the production of autophagosomal membranes directly or indirectly through their specific structures to alter membrane curvature (Liu et al. 2023). There are three common mechanisms to explain the change in membrane curvature. For example, the BAR domain of Bif-1 senses and tethers Atg9 vesicles to change the membrane curvature of the isolation membrane (IM), and the Atg9 vesicles are reported as a source of the IM in the autophagy process. The amphiphilic helix of Bif-1 inserts directly into the phospholipid bilayer, causing membrane asymmetry, and thus changing the membrane curvature of the IM. Atg2 forms a pathway for lipid transport from the endoplasmic reticulum to the IM, and this pathway also contributes to the formation of the IM. Liu et al. 2023 introduced the phenomena and causes of membrane curvature changes in the process of macroautophagy as well as the mechanisms of ATGs in membrane curvature and autophagosome membrane formation.
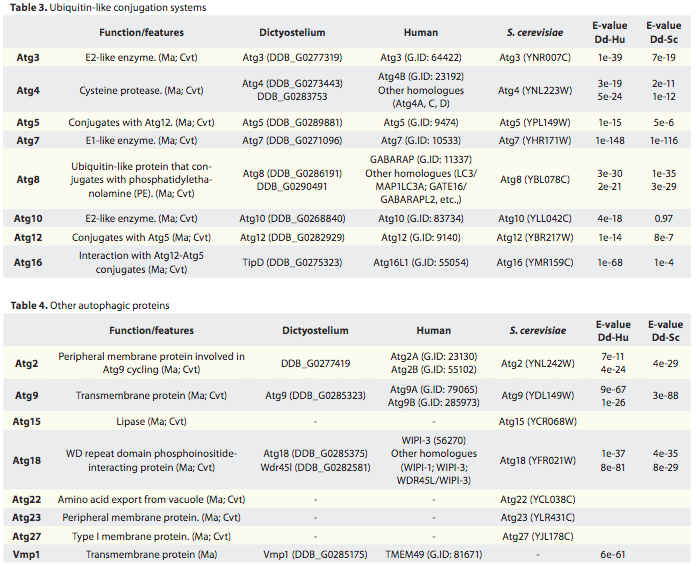
The table (Table 3) presented below is taken from (Munakata and Klionsky 2010).